All content on this site is intended for healthcare professionals only. By acknowledging this message and accessing the information on this website you are confirming that you are a healthcare professional. If you are a patient or carer, please visit the MPN Advocates Network.
The MPN Hub website uses a third-party service provided by Google that dynamically translates web content. Translations are machine generated, so may not be an exact or complete translation, and the MPN Hub cannot guarantee the accuracy of translated content. The MPN Hub and its employees will not be liable for any direct, indirect, or consequential damages (even if foreseeable) resulting from use of the Google Translate feature. For further support with Google Translate, visit Google Translate Help.
The MPN Hub is an independent medical education platform, sponsored by AOP Health, GSK, and Novartis, and supported through independent educational grants from Blueprint Medicines, Incyte, Bristol Myers Squibb and Merck Sharp & Dohme LLC, Rahway, NJ, USA. The funders are allowed no direct influence on our content. The levels of sponsorship listed are reflective of the amount of funding given. View funders.
Now you can support HCPs in making informed decisions for their patients
Your contribution helps us continuously deliver expertly curated content to HCPs worldwide. You will also have the opportunity to make a content suggestion for consideration and receive updates on the impact contributions are making to our content.
Find out more
Create an account and access these new features:
Bookmark content to read later
Select your specific areas of interest
View MPN content recommended for you
Molecular mechanisms associated with leukemic transformation of PV and ET
Leukemic transformation in patients with polycythemia vera (PV) and essential thrombocythemia (ET) is considered a leading long-term complication, occurring in 2─5% of cases at 15 years. Patients who develop secondary acute myeloid leukemia (AML) have a poor prognosis, and the identification of PV and ET patients at high risk of leukemic transformation would help in monitoring and treating these patients, however it remains difficult.
Mutations in TP53, ASXL1, RUNX1, EZH2, IDH1/2, and SH2B3 are associated with high risk of leukemic transformation, but little is known about predictors of time to leukemic evolution. To better understand genetic mechanisms associated with leukemia development, Damien Luque Paz and colleagues, in a study published in Blood Advances, performed molecular analyses at different time points during disease evolution (at diagnosis, during the chronic phase, and at transformation) in patients with PV or ET who developed AML.1
Study design
Patients included in the study had previously been diagnosed with PV (n = 24) or ET (n = 25) and showed leukemic evolution. Molecular analyses were performed by next-generation sequencing on DNA samples obtained at the time of leukemic evolution and at diagnosis, or at the chronic phase (the earliest available sample was used if the sample at diagnosis was unavailable; Figure 1). In addition, a control cohort of 80 patients with PV (n = 37) or ET (n = 43) who did not develop AML after ≥ 8 years of follow-up was studied to investigate the prognosis of additional mutations during the chronic phase.
Figure 1. Cohort description1
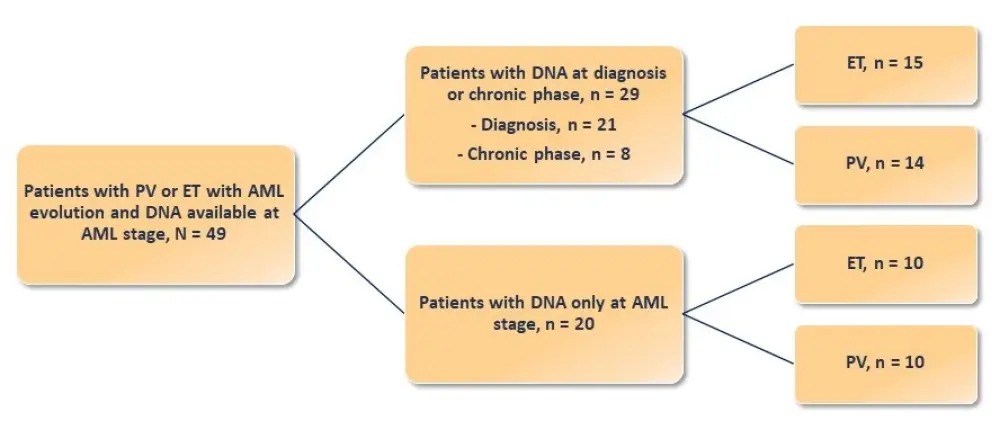
Patient characteristics
The characteristics of 49 patients with PV or ET who developed AML are shown in Table 1.
Table 1. Patient characteristics1
|
ET, essential thrombocythemia; PV, polycythemia vera. |
|||
|
Characteristic |
Entire cohort |
PV |
ET |
|---|---|---|---|
|
Median age at diagnosis, years (range) |
63 (26─82) |
65 (42─81) |
59 (26─82) |
|
Hemoglobin at diagnosis, g/dL (range) |
16 (10─20.7) |
18.1 (16─20.7) |
14.4 (10─16.6) |
|
Platelets at diagnosis, ×109/L (range) |
624 (139─1,575) |
404 (139─748) |
800 (471─1,575) |
|
Leukocytes at diagnosis, ×109/L (range) |
9.17 (3─27.8) |
10.6 (5.1─27.8) |
8.4 (3─17.3) |
|
Driver mutation, n (%) |
|
|
|
|
JAK2V617F |
39 (80) |
24 (100) |
15 (60) |
|
CALR |
6 (12) |
— |
6 (24) |
|
MPL |
2 (4) |
— |
2 (8) |
|
Triple negative |
2 (4) |
— |
2 (8) |
|
Time to leukemic transformation, years (range) |
12 (1─30) |
11 (2─30) |
12 (1─30) |
|
Karyotype at leukemic transformation, n (%) |
n = 35 |
n = 17 |
n = 18 |
|
Normal |
13 (37) |
6 (35) |
7 (39) |
|
Abnormal not complex |
6 (17) |
4 (24) |
2 (11) |
|
Complex |
9 (26) |
5 (29) |
4 (22) |
|
Monosomal |
7 (20) |
2 (12) |
5 (28) |
Results
The total number of additional mutations (in addition to driver mutations) detected in the entire cohort was 191, with the most frequently mutated genes being TP53, TET2, RUNX1, ASXL1, and EZH2. At least one additional mutation was present in all patients with a median number of four mutations. The time to leukemic transformation was variable among patients and seemed to be related to the genes affected by additional mutations. Through sparse partial least squares discriminant analysis and using a hierarchical classification, the authors identified three groups of patients with different time to leukemic evolution (Figure 2):
- Short-term (median time to transformation: 3 years) group presented a complex mutational landscape with preferential mutations in RUNX1, IDH1/2, U2AF1, and TET2 genes.
- Intermediate (median time to transformation: 10 years) and long-term (median time to transformation: 21 years) groups presented a less complex molecular landscape, with mutations occurring in the TP53 and NRAS genes.
Figure 2. Molecular landscape1
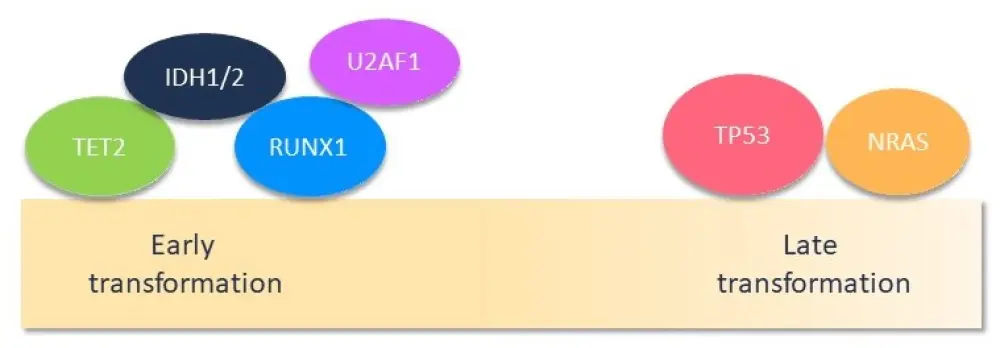
To better understand the mutations and allele burden evolution between the chronic and leukemic phases, matched samples obtained at diagnosis or during the chronic phase, available for 29 patients (Figure 1), were evaluated:
- In 8 out of 23 (35%) cases, the driver mutation JAK2V617F disappeared at transformation.
- IDH1/2, TET2, DNMT3A, and splicing (SRSF2, U2AF1, and ZRSR2) genes were frequently mutated at diagnosis (68% of cases).
- Mutations in NRAS and RUNX1 were rarely detected at diagnosis.
- Mutations in TP53, RUNX1, BCOR, EZH2, and ASXL1 were present at a higher allelic burden during the period of leukemic transformation.
In addition, samples between diagnosis and leukemic transformation were analyzed in seven patients:
- In five patients, several mutations observed at transformation were not detectable 1─2 years earlier, especially in the TP53 and RUNX1 genes.
- In one patient, mutations in ASXL1, EZH2, and NRAS were detectable 1 year before evolution to AML.
- TP53 mutations were late events.
Impact of additional mutations during the chronic phase on leukemia-free survival
To evaluate the impact of additional mutations during the chronic phase on the risk of transformation, 29 patients with post-PV or -ET AML with a sample obtained during the chronic phase were compared with 80 stable patients with myeloproliferative neoplasms (PV, n = 37; ET, n = 43) who did not develop AML during ≥ 8 years of follow-up (range, 8─29 years):
- In the control cohort
- additional mutations were detected in 28 cases (35%); and
- TET2 was the most mutated gene (11 patients with ≥ 1 mutation [14%]).
- The presence of ≥ 1 additional mutation during the chronic phase was associated with a higher risk of leukemic transformation (p < 0.0001); a stronger association was observed after the exclusion of TET2 mutations from the analysis (HR, 7.3; 95% CI, 2.9-18; p < 0.0001).
- In multivariate analysis, both age at the time of sampling (HR, 1.07; 95% CI, 1.03─1.12; p = 0.0004) and the presence of ≥ 1 additional mutation (HR, 4.8; 95% CI, 1.8─13; p = 0.002) were associated with a worse leukemia-free survival.
- The median overall survival (OS) after transformation in the short-term, intermediate, and long-term groups was 19, 4, and 8 months, respectively.
- The 18-month OS with best supportive care was 9% and was similar to intensive therapy. By contrast, the 18-month OS for nonintensive therapy, and hematopoietic stem cell transplantation was 15%, and 50%, respectively (p = 0.015).
Prognostic impact of additional mutations
The prognostic impact of additional mutations in TET2, ASXL1, RUNX1, and TP53 was examined in ten patients at leukemic transformation. Patients with mutated TP53 had a worse OS compared with patients with wild-type TP53 (12-month OS, 18% vs 48%, respectively; p = 0.05).
Conclusion
In this study, the authors evaluated not only the presence/absence of mutations at leukemic transformation, but also the allele burden. The findings suggested that mutations that show an increasing allele burden are likely to contribute to the risk of transformation to AML. They identified three molecular groups associated with different time to transformation:
- Short-term transformation group characterized by mutations in IDH1/2, RUNX1, and U2AF1 genes.
- Intermediate and long-term transformation groups associated with mutations in TP53, NRAS, and BCORL1 genes.
Some of these mutations were already present at diagnosis/chronic phase, either with a significant allele burden or with a very low allele burden (especially TP53 mutations). An allele burden > 50% or a second mutation involving TP53 was observed in most leukemic transformation events, which may explain the inferior outcome to intensive therapy. Taken together these results suggest that the leukemic transformation of PV and ET involves different time-dependent molecular mechanisms, however further functional studies and larger cohorts are required to better understand these mechanisms.
References
Please indicate your level of agreement with the following statements:
The content was clear and easy to understand
The content addressed the learning objectives
The content was relevant to my practice
I will change my clinical practice as a result of this content

